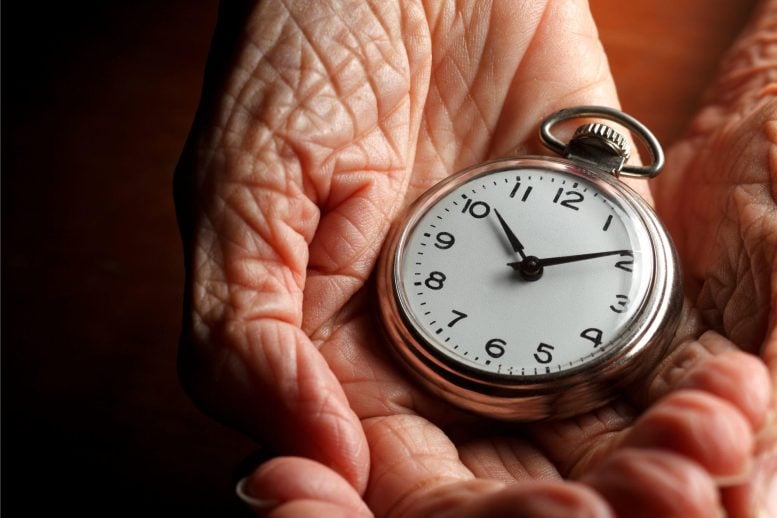

Aging clocks, which measure biological age with precision, can deviate from chronological age due to environmental influences like smoking or diet. Researchers at the University of Cologne found that these clocks actually track increasing random cellular changes, suggesting that biological aging could be influenced by stochastic variations in processes like DNA methylation and gene activity.
Aging clocks can accurately determine a person’s biological age, which can differ from their chronological age—the age calculated from their date of birth—due to environmental influences like diet or smoking. The precision of these clocks indicates that the aging process follows a program.
Scientists David Meyer and Professor Dr. Björn Schumacher at CECAD, the Cluster of Excellence Cellular Stress Responses in Aging-Associated Diseases of the University of Cologne, have now discovered that aging clocks actually measure the increase in stochastic changes in cells. The study was recently published published in Nature Aging.
“Aging is triggered when the building blocks in our cells become damaged. Where this damage occurs is for the most part random. Our work combines the accuracy of aging clocks with the accumulation of entirely stochastic changes in our cells,” said Professor Schumacher.
Fewer Checks, More Noise
With increasing age, controlling the processes that occur in our cells becomes less effective, resulting in more stochastic results. This is particularly evident in the accumulation of stochastic changes in DNA methylation. Methylation refers to the chemical changes that affect DNA, the genome’s building blocks. These methylation processes are strictly regulated within the body. However, during the course of one’s life, random changes occur in the methylation patterns. The accumulation of variation is a highly accurate indicator of a person’s age.
The loss of control over the cells and the increase in stochastic variation are not restricted to DNA methylation. Meyer and Schumacher demonstrate that the increase in stochastic variations also in gene activity can be used as an aging clock. “In principle it would be feasible to take this even further, allowing the stochastic variations in any process in the cell to predict age,” Schumacher said. According to the authors, it is above all crucial to ascertain if such aging clocks can show the success of interventions that slow the aging process or harmful factors that accelerate aging.
Using the available datasets, the scientists showed that smoking increases the random changes in humans and that ‘anti-aging’ interventions such as lower calorie intake in mice reduce the variation in methylation patterns. They also showed that the stochastic noise is even reversible by means of reprogramming body cells to stem cells. The scientists compared human fibroblasts from the skin that were reprogrammed into stem cells and as a result of the reprogramming are rejuvenating. The high variation indicative of the age of the body cells was indeed reversed to the low stochastic noise of young stem cells.
Meyer and Schumacher hope that their findings on the loss of regulation and the accumulating stochastic variations will lead to new interventions that can tackle the root cause of aging and may even lead to cellular rejuvenation. A target for such interventions could be repairing stochastic changes in DNA or improved control of gene expression.
Reference: “Aging clocks based on accumulating stochastic variation” by David H. Meyer, and Björn Schumacher, 9 May 2024, Nature Aging.
DOI: 10.1038/s43587-024-00619-x







From the article: “Using the available datasets, the scientists showed that smoking increases the random changes in humans and that ‘anti-aging’ interventions such as lower calorie intake in mice reduce the variation in methylation patterns.”
Higher education seems to blind researchers to the “…forest for the trees.” For a start, as a now eighty year old American male who’s been compelled by officially (FDA in the US) approved food poisoning to explore new areas of the ‘forest’ for forty-three years and counting with some successes, I can state with some confidence, again, that the “…available datasets…” have multiple fatal flaws. First, they don’t factor-in a kind of nearly subclinical non-IgE-mediated food (minimally; other) allergy reaction which can be acquired and/or inherited. More specifically, when it comes to ‘smoking,’ a person’s individual allergic reaction to tobacco or some substance in the smoke is probably what makes some smokers ill and others not. Next, they don’t know to factor-in the relationship between repetitious trauma and acquired allergies; a psychosomatic factor. Additionally, the ‘available datasets’ do not factor-in increasing numbers of officially approved toxic food additives, namely added soy since the late 1960s, MSG since 1980 and the increasingly pervasive cooking oil preservative TBHQ since 1972; others. Finally, for now, they don’t yet factor-in allergy related acidic blood and/or possible nutritional deficiencies like calcium (unreliable serum testing), especially in mothers due to the calcium demands of the fetus, and they certainly don’t factor-in how it is probably where the acidic flow of blood is the greatest that the greatest immediate physical harm and ‘DNA methylation’ occur.
Cell rejuvenation aside, as long as the primary goal of researchers is to find ‘new interventions’ instead of identifying and eliminating ‘root causes,’ the ‘forest’ is likely to remain invisibly hidden behind huge trees of deceit, medical errors, ignorance and fatally flawed so-called “evidence-based” datasets.